Simulations¶
The LiteBIRD Simulation Framework is built on the Simulation
class, which should be instantiated in any pipeline built using this
framework. The class acts as a container for the many analysis modules
available to the user, and it offers the following features:
Provenance model;
Interface with the instrument database;
System abstractions;
Generation of reports;
Printing status messages on the terminal (logging).
Provenance model¶
A «provenance model» is, generally speaking, a way to track the history and origin of a data set by recording the following information:
Who or what created the dataset?
Which algorithm or instrumentation was used to produce it?
Which steps were undertaken to process the raw data?
How can one get access to the raw samples used to produce the dataset?
The LiteBIRD Simulation Framework tracks these information using parameter files (in TOML format), accessing the LiteBIRD Instrument Model (IMO) through the unique identifiers provided by the Data Access Layer (DAL), and generating reports after the simulation completes.
Parameter files¶
When you run a simulation, there are typically plenty of parameters that need to be passed to the code: the resolution of an output map, the names of the detectors to simulate, whether to include synchrotron emission in the sky model, etc.
The Simulation class eases this task by accepting the path to
a TOML file as a parameter (parameter_file). Specifying this
parameter triggers two actions:
The file is copied to the output directory where the simulation output files are going to be written (i.e., the same that contains
report.html);The file is read and made available in the field
Simulation.parameters(a Python dictionary).
Using a parameter file is optional; if you do not specify
parameter_file while creating a Simulation object, the
parameters field will be set to an empty dictionary. (You can even
directly pass a dictionary to a Simulation object: this can
be handy if you already constructed a parameter object somewhere
else.)
Take this example of a simple TOML file:
# This is file "my_conf.toml"
[simulation]
random_seed = 12345
[general]
nside = 512
imo_version = "v1.3"
[sky_model]
components = ["synchrotron", "dust", "cmb"]
The following example loads the TOML file and prints its contents to the terminal:
import litebird_sim as lbs
sim = lbs.Simulation(parameter_file="my_conf.toml")
print("Seed for the random number generator:",
sim.parameters["simulation"]["random_seed"])
print("NSIDE =", sim.parameters["general"]["nside"])
print("The IMO I'm going to use is",
sim.parameters["general"]["imo_version"])
print("Here are the sky components I'm going to simulate:")
for component in sim.parameters["sky_model"]["components"]:
print("-", component)
The output of the script is the following:
Seed for the random number generator: 12345
NSIDE = 512
The IMO I'm going to use is v1.3
Here are the sky components I'm going to simulate:
- synchrotron
- dust
- cmb
A Simulation object only interprets the section
simulation and leaves everything else unevaluated: it’s up to the
simulation modules to make sense of any other section. The recognized
parameters in the section named simulation are the following:
base_path: a string containing the path where to save the results of the simulation.start_time: the start time of the simulation. If it is a string or a TOML datetime, it will be passed to the constructor forastropy.time.Time, otherwise it must be a floating-point value.duration_s: a floating-point number specifying how many seconds the simulation should last. You can pass a string, which can contain a measurement unit as well: in this case, you are not forced to specify the duration in seconds. Valid units are:days(orday),hours(orhour),min, andsec(ors).name: a string containing the name of the simulation.description: a string containing a (possibly long) description of what the simulation does.random_seed: the seed for the random number generator. An integer value ensures the reproducibility of the results obtained with random numbers; by passingNonethere will not be the possibility to re-obtain the same outputs. You can find more details in Random numbers in litebird_sim.numba_num_of_threads: number of threads used by Numba when running a parallel calculation.numba_threading_layer: the multi-threading library to be used by Numba for parallel computations. See the Numba user’s manual to check which choices are available.
These parameters can be used instead of the keywords in the
constructor of the Simulation class. Consider the following
code:
sim = lbs.Simulation(
base_path="/storage/output",
start_time=astropy.time.Time("2020-02-01T10:30:00"),
duration_s=3600.0,
name="My simulation",
description="A long description should be put here",
random_seed=12345,
numba_num_of_threads=4,
numba_threading_layer="workqueue",
)
You can achieve the same if you create a TOML file named foo.toml
that contains the following lines:
[simulation]
base_path = "/storage/output"
start_time = 2020-02-01T10:30:00
duration_s = 3600.0
name = "My simulation"
description = "A long description should be put here"
random_seed = 12345
numba_num_of_threads = 4
numba_threading_layer = "workqueue"
and then you initialize the sim variable in your Python code as follows:
sim = lbs.Simulation(parameter_file="foo.toml")
A nice feature of the framework is that the duration of the mission must not be specified in seconds. You would achieve identical results if you specify the duration in one of the following ways:
# All of these are the same
duration_s = "1 hour"
duration_s = "60 min"
duration_s = "3600 s"
Interface with the instrument database¶
To simulation LiteBIRD’s data acquisition, the simulation code must be aware of the characteristics of the instrument. These are specified in the LiteBIRD Instrument Model (IMO) database, which can be accessed by people with sufficient rights. This Simulation Framework has the ability to access the database and take the input parameters necessary for its analysis modules to produce the expected output.
System abstractions¶
In some cases, simulations must be ran on HPC computers, distributing the job on many processing units; in other cases, a simple laptop might be enough. The LiteBIRD Simulation Framework uses MPI to parallelize its codes, which is however an optional dependency: the code can be ran serially.
If you want to use MPI in your scripts, assuming that you have created a virtual environment as explained in the Tutorial, you have just to install mpi4py:
$ pip install mpi4py
# Always do this after "pip install", it will record the
# version number of mpi4py and all the other libraries
# you have installed in your virtual environment
$ pip freeze > requirements.txt
# Run the program using 4 processes
$ mpiexec -n 4 python3 my_script.py
The framework will detect the presence of mpi4py, and it will
automatically use distributed algorithms where applicable; otherwise,
serial algorithms will be used. You can configure this while creating
a Simulation object:
import litebird_sim as lbs
import mpi4py
# This simulation *must* be ran using MPI
sim = lbs.Simulation(mpi_comm=mpi4py.MPI.COMM_WORLD, random_seed=12345)
The framework sets a number of variables related to MPI; these
variables are always defined, even if MPI is not available, and they
can be used to make the code work in serial environments too. If your
code must be able to run both with and without MPI, you should
initialize a Simulation object using the variable
MPI_COMM_WORLD, which is either mpi4py.MPI.COMM_WORLD
(the default MPI communicator) or a dummy class if MPI is disabled:
import litebird_sim as lbs
# This simulation can take advantage of MPI if present,
# otherwise it will stick to serial execution
sim = lbs.Simulation(mpi_comm=lbs.MPI_COMM_WORLD, random_seed=12345)
See the page Multithreading and MPI for more information.
Generation of reports¶
This section should explain how reports can be generated, first from the perspective of a library user, and then describing how developers can generate plots for their own modules.
Here is an example, showing several advanced topics like mathematical formulae, plots, and value substitution:
import litebird_sim as lbs
import matplotlib.pylab as plt
sim = lbs.Simulation(
name="My simulation",
base_path="output",
random_seed=12345,
)
data_points = [0, 1, 2, 3]
plt.plot(data_points)
fig = plt.gcf()
sim.append_to_report('''
Here is a formula for $`f(x)`$:
```math
f(x) = \sin x
```
And here is a completely unrelated plot:

The data points have the following values:
{% for sample in data_points %}
- {{ sample }}
{% endfor %}
''', figures=[(fig, "myplot.png")],
data_points=data_points)
sim.flush()
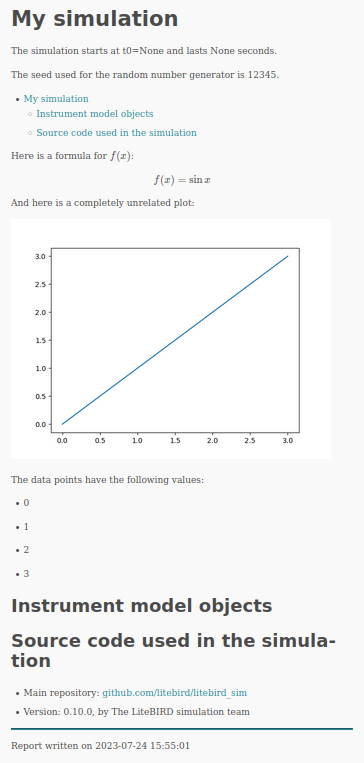
And here is the output, which is saved in output/report.html:
Logging¶
The report generation tools described above are useful to produce a
synthetic report of the scientific outcomes of a simulation.
However, one often wants to monitor the execution of the code in a
more detailed manner, checking which functions have been called, how
often, etc. In this case, the best option is to write messages to the
terminal. Python provides the logging module for this
purpose, and when you initialize a Simulation object, the
module is initialized with a set of sensible defaults. In your code you
can use the functions debug, info, warning, error, and
critical to monitor what’s happening during execution:
import litebird_sim as lbs
import logging as log # "log" is shorter to write
my_sim = lbs.Simulation(random_seed=12345)
log.info("the simulation starts here!")
pi = 3.15
if pi != 3.14:
log.error("wrong value of pi!")
The output of the code above is the following:
[2020-07-18 06:25:27,653 INFO] the simulation starts here!
[2020-07-18 06:25:27,653 ERROR] wrong value of pi!
Note that the messages are prepended with the date, time, and level of severity of the message.
A few environment variables can taylor the way logging is done:
LOG_DEBUG: by default, debug messages are not printed to the terminal, because they are often too verbose for typical uses. If you want to debug your code, set a non-empty value to this variable.LOG_ALL_MPI: by default, if you are using MPI then only messages from the process running with rank 0 will be printed. Setting this environment variable will make all the processes print their message to the terminal. (Caution: there might be overlapping messages, if two processes happen to write at the same time.)
The way you use these variable from the terminal is illustrated with
an example. Suppose that we changed our example above, so that
log.debug is called instead of log.info:
import litebird_sim as lbs
import logging as log # "log" is shorter to write
my_sim = lbs.Simulation(random_seed=12345)
log.debug("the simulation starts here!")
pi = 3.15
if pi != 3.14:
log.debug("wrong value of pi!")
In this case, running the script will produce no messages, as the
default is to skip log.debug calls:
$ python my_script.py
$
However, running the script with the environment variable
LOG_DEBUG set will make the messages appear:
$ LOG_DEBUG=1 python my_script.py
[2020-07-18 06:31:03,223 DEBUG] the simulation starts here!
[2020-07-18 06:31:03,224 DEBUG] wrong value of pi!
$
Monitoring MPI processes¶
When using MPI, the method Simulation.create_observations()
distributes detectors and time spans over all the available
MPI processes. The Simulation class provides a method
that enables the user to inspect how the TOD has been split
among the many MPI processes: Simulation.describe_mpi_distribution().
The method must be called at the same time on all the MPI
processess, once you have successfully called
Simulation.create_observations():
import litebird_sim as lbs
# Be sure to create a `Simulation` object and create
# the observations:
#
# sim = lbs.Simulation(…)
# sim.create_observations(…)
# Now ask the framework to report how it's using each MPI process
descr = sim.describe_mpi_distribution()
The return type for Simulation.describe_mpi_distribution()
is MpiDistributionDescr, which can be printed on the
spot using print(descr). The result looks like the following
example:
# MPI rank #1
## Observation #0
- Start time: 0.0
- Duration: 21600.0 s
- 1 detector(s) (0A)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
## Observation #1
- Start time: 43200.0
- Duration: 21600.0 s
- 1 detector(s) (0A)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
# MPI rank #2
## Observation #0
- Start time: 21600.0
- Duration: 21600.0 s
- 1 detector(s) (0A)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
## Observation #1
- Start time: 64800.0
- Duration: 21600.0 s
- 1 detector(s) (0A)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
# MPI rank #3
## Observation #0
- Start time: 0.0
- Duration: 21600.0 s
- 1 detector(s) (0B)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
## Observation #1
- Start time: 43200.0
- Duration: 21600.0 s
- 1 detector(s) (0B)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
# MPI rank #4
## Observation #0
- Start time: 21600.0
- Duration: 21600.0 s
- 1 detector(s) (0B)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
## Observation #1
- Start time: 64800.0
- Duration: 21600.0 s
- 1 detector(s) (0B)
- TOD(s): tod
- TOD shape: 1×216000
- Types of the TODs: float64
The class MpiDistributionDescr contains a list
of MpiProcessDescr, which describe the «contents»
of each MPI process. The most important field in each
MpiProcessDescr instance is observations, which
is a list of MpiObservationDescr objects: it
contains the size of the TOD array, the names of the detectors,
and other useful information. Refer to the documentation of
each class to know what is inside.
High level interface¶
The class Simulation has a powerful high level
interface that allows to quickly generate a scanning strategy,
allocate the observations, generate simulated timelines
cointaing signal, noise and dipole, build maps, and
save(read) the entire simulation object. The syntax is
straightforward:
sim = lbs.Simulation(...)
sim.set_scanning_strategy(...)
sim.set_instrument(...)
sim.create_observations(...)
sim.compute_pointings()
sim.compute_pos_and_vel()
sim.add_dipole()
sim.fill_tods(...)
sim.add_noise(...)
result = sim.make_destriped_map(nside=nside)
healpy.mollview(result.destriped_map)
sim.write_observations(...)
sim.read_observations(...)
See the documentation in Observations, Scanning strategy Dipole anisotropy, Time Ordered Simulations, Map-making for details of the single functions.
Data splits¶
Since the class Simulation is interfaced to
litebird_sim.make_binned_map(), it is able to provide data splits
both in time and detector space (see Map-making
for more details on the splitting and the available options).
In addition, the class contains Simulation.make_binned_map_splits(),
which is a wrapper around litebird_sim.make_binned_map()
that allows to perform the mapmaking on multiple choices
of splits at once (passed as a list of strings).
Indeed, the function will loop over the cartesian
product of the time and detector splits. The default
behavior is to perform the mapmaking in each combination
and save the result to disk, to avoid memory issues. In particular,
only the binned maps are saved to disk, unless you set
include_inv_covariance to True. This saves the elements of
the inverse covariance as extra fields in the output FITS file.
This default behavior can be changed by setting the write_to_disk parameter
to False. Then, the function returns a dictionary
containing the full results of the mapmaking for each split.
The keys are strings that describe the split in the
format “{Dsplit}_{Tsplit}”, such as “waferL00_year1”.
Before performing the computation, the function Simulation.check_valid_splits()
will check whether the requested split is valid (see Map-making).
This is a wrapper around litebird_sim.check_valid_splits(). If the
split is not valid, the function will raise a ValueError. In addition, it
will check whether the requested split is compatible with the duration of
the observation and with the detector list. Thus, for example, if the
observation lasts 1 year, the split “year2” will raise an AsserionError. Similarly,
if the observation is performed with some detector contained in the L00
wafer, the split “waferL03” will also raise an AsserionError.
Profiling a simulation¶
The class Simulation provides an higher-level interface to the
functions described in the chapter Profiling. The decorator @profile
automatically wraps a method of the Simulation class so that it measures
the time spent to run the method and adds it to an internal list of
TimeProfiler objects. When the Simulation.flush() method is
called, each MPI process creates its own JSON file containing the methods that
have been measured during the execution of the simulation. These files are
stored in the output directory and have names like profile_mpi00000.json,
profile_mpi00001.json, etc.
API reference¶
- class litebird_sim.simulations.MpiDistributionDescr(num_of_observations: int, detectors: List[DetectorInfo], mpi_processes: List[MpiProcessDescr])¶
Bases:
objectA class that describes how observations are distributed among MPI processes
The fields defined in this dataclass are the following:
num_of_observations (int): overall number of observations in all the MPI processes
detectors (list of
DetectorInfoobjects): list of all the detectors used in the observationsmpi_processes: list of
MpiProcessDescrinstances, describing the kind of data that each MPI process is currently holding
Use
Simulation.describe_mpi_distribution()to get an instance of this object.- detectors: List[DetectorInfo]¶
- mpi_processes: List[MpiProcessDescr]¶
- num_of_observations: int¶
- class litebird_sim.simulations.MpiObservationDescr(det_names: List[str], tod_names: List[str], tod_shape: Tuple[int, int] | None, tod_dtype: List[str], tod_description: List[str], start_time: float | Time, duration_s: float, num_of_samples: int, num_of_detectors: int)¶
Bases:
objectThis class is used within
MpiProcessDescr. It describes the kind and size of the data held by aObservationobject.Its fields are:
det_names (list of
str): names of the detectors handled by this observationtod_names (list of
str): names of the fields containing the TODs (e.g.,tod,cmb_tod,dipole_tod, …)tod_shape (tuples of
int): shape of each TOD held by the observation. This is not a list, because all the TODs are assumed to have the same shapetod_dtype (list of
str): string representing the NumPy data type of each TODs, in the same order as in the field tod_nametod_description (list of
str): list of human-readable descriptions for each TOD, in the same order as in the field tod_namestart_time (either a
floator aastropy.time.Time): start date of the observationduration_s (
float): duration of the TOD in secondsnum_of_samples (
int): number of samples held by this TODnum_of_detectors (
int): number of detectors held by this TOD. It’s the length of the field det_names (see above)
- det_names: List[str]¶
- duration_s: float¶
- num_of_detectors: int¶
- num_of_samples: int¶
- start_time: float | Time¶
- tod_description: List[str]¶
- tod_dtype: List[str]¶
- tod_names: List[str]¶
- tod_shape: Tuple[int, int] | None¶
- class litebird_sim.simulations.MpiProcessDescr(mpi_rank: int, numba_num_of_threads: int, observations: List[MpiObservationDescr])¶
Bases:
objectDescription of the kind of data held by a MPI process
This class is used within
MpiDistributionDescr. Its fields are:mpi_rank: rank of the MPI process described by this instance
observations: list of
MpiObservationDescrobjects, each describing one observation managed by the MPI process with rank mpi_rank.- numba_num_of_threads (
int): number of threads used by Numba in the MPI process with rank mpi_rank.
- numba_num_of_threads (
- mpi_rank: int¶
- numba_num_of_threads: int¶
- observations: List[MpiObservationDescr]¶
- class litebird_sim.simulations.OutputFileRecord(path, description)¶
Bases:
tuple- description¶
Alias for field number 1
- path¶
Alias for field number 0
- class litebird_sim.simulations.Simulation(random_seed='', base_path=None, name=None, mpi_comm=<litebird_sim.mpi._SerialMpiCommunicator object>, description='', start_time=None, duration_s=None, imo=None, parameter_file=None, parameters=None, numba_threads=None, numba_threading_layer=None, profile_time: bool = True)¶
Bases:
objectA container object for running simulations
This is the most important class in the Litebird_sim framework. It initializes an output directory that will contain all the products of a simulation and will handle the generation of reports and writing of output files.
Be sure to call
Simulation.flush()when the simulation is completed. This ensures that all the information are saved to disk before the completion of your script.You can access the fields base_path, name, mpi_comm, and description in the Simulation object:
sim = litebird_sim.Simulation(name="My simulation") print(f"Running {sim.name}, saving results in {sim.base_path}")
The member variable observations is a list of
Observationobjects, which is initialized by the methodcreate_observations().This class keeps track of any output file saved in base_path through the member variable self.list_of_outputs. This is a list of objects of type
OutputFileRecord(), which are 2-tuples of the form(path, description), wherepathis apathlib.Pathobject anddescriptionis a str object:for curpath, curdescr in sim.list_of_outputs: print(f"{curpath}: {curdescr}")
When pointing information is needed, you can call the method
Simulation.set_scanning_strategy(), which initializes the members pointing_freq_hz and spin2ecliptic_quats; these members are used by functions likeget_pointings().- Parameters:
random_seed (int or None) – the seed used for the random number generator. The user is required to set this parameter. By setting it to None, the generation of random numbers will not be reproducible.
base_path (str or pathlib.Path) – the folder that will contain the output. If this folder does not exist and the user has sufficient rights, it will be created.
name (str) – a string identifying the simulation. This will be used in the reports.
mpi_comm – either None (do not use MPI) or a MPI communicator object, like mpi4py.MPI.COMM_WORLD.
description (str) – a (possibly long) description of the simulation, to be put in the report saved in base_path).
start_time (float or
astropy.time.Time) – the start time of the simulation. It can be either an arbitrary floating-point number (e.g., 0) or anastropy.time.Timeinstance; in the latter case, this triggers a more precise (and slower) computation of pointing information.duration_s (float) – Number of seconds the simulation should last.
imo (
Imo) – an instance of theImoclass. If not provided, the default constructor will be called, which means that the IMO to be used will be the default one configured using theinstall_imoprogram.parameter_file (str or pathlib.Path) – path to a TOML file that contains the parameters for the simulation. This file will be copied into base_path, and its contents will be read into the field parameters (a Python dictionary).
numba_threads (int) – number of threads to use when calling Numba functions. If no value is passed but the environment variable
OMP_NUM_THREADSis set, its value will be used; otherwise a number of threads equal to the number of CPU cores will be used.numba_threading_layer (str) – name of the Numba threading layer to use. See the Numba User’s Manual: <https://numba.readthedocs.io/en/stable/user/threading-layer.html>
profile_time (bool) – if
True(the default), record information about the time spent while doing some time-consuming tasks.
- add_dipole(**kwargs)¶
- add_noise(**kwargs)¶
- append_to_report(markdown_text: str, append_newline=True, figures: List[Tuple[Any, str]] = [], **kwargs)¶
Append text and figures to the simulation report
- Parameters:
markdown_text (str) – text to be appended to the report.
append_newline (bool) – append newlines to the end of the text. This ensures that calling again this method will produce a separate paragraph.
figures (list of 2-tuples) – list of Matplotlib figures to be saved in the report. Each tuple must contain one figure and one filename. The figures will be saved using the specified file name in the output directory. The file name must match the one used as reference in the Markdown text.
kwargs – any other keyword argument will be used to expand the text markdown_text using the Jinja2 library library.
A Simulation class can generate reports in Markdown format. Use this function to add some text to the report, possibly including figures. The function has no effect if called from an MPI rank different from #0.
It is possible to use objects other than Matplotlib figures. The only method this function calls is savefig, with no arguments.
Images are saved immediately during the call, but the text will be written to disk only when
flush()is called.You can put LaTeX formulae in the text, using
$`...`$for inline equations and the math tag in fenced text for displayed equations.
- apply_gaindrift(**kwargs)¶
- check_valid_splits(detector_splits, time_splits)¶
Wrapper around
litebird_sim.check_valid_splits(). Checks that the splits are valid on the observations.
- compute_pointings(**kwargs)¶
- compute_pos_and_vel(**kwargs)¶
- create_observations(**kwargs)¶
- describe_mpi_distribution() MpiDistributionDescr | None¶
Return a
MpiDistributionDescrobject describing observationsThis method returns a
MpiDistributionDescrthat describes the data stored in each MPI process running concurrently. It is a great debugging tool when you are using MPI, and it can be used for tasks where you have to carefully orchestrate they way different MPI processes run together.If this method is called before
Simulation.create_observations(), it will returnNone.This method should be called by all the MPI processes. It can be executed in a serial environment (i.e., without MPI) and will still return meaningful values.
The typical usage for this method is to call it once you have called
Simulation.create_observations()to check that the TODs have been laid in memory in the way you expect:sim.create_observations(…) distr = sim.describe_mpi_distribution() if litebird_sim.MPI_COMM_WORLD.rank == 0: print(distr)
- distribute_workload(observations: List[Observation])¶
- fill_tods(**kwargs)¶
- flush(include_git_diff=True, base_imo_url: str = 'https://litebirdimo.ssdc.asi.it', profile_file_name: str | None = None)¶
Terminate a simulation.
This function must be called when a simulation is complete. It will save pending data to the output directory.
It returns a Path object pointing to the HTML file that has been saved in the directory pointed by
self.base_path.
- generate_spin2ecl_quaternions(scanning_strategy: None | ScanningStrategy = None, imo_url: None | str = None, delta_time_s: float = 60.0, append_to_report=True)¶
Deprecated since version 0.9: Use set_scanning_strategy
- get_list_of_tods() List[TodDescription]¶
- get_tod_descriptions() List[str]¶
- get_tod_dtypes() List[Any]¶
- get_tod_names() List[str]¶
- init_random(random_seed)¶
Initialize a random number generator in the random field
This function creates a random number generator and saves it in the field random. It should be used whenever a random number generator is needed in the simulation. In the case random_seed has not been set to None, it ensures that different MPI processes have their own different seed, which stems from the parameter random_seed, and the results will be reproducible. The generator is PCG64, and it is ensured that the sequences in each MPI process are independent. If init_random is called with the same seed but a different number of MPI ranks, the sequence of random numbers will be different. In the case random_seed has been set to None, no seed will be used and the results obtained with the random number generator will not be reproducible.
This method is automatically called in the constructor, but it can be called again as many times as required. The typical case is when one wants to use a seed that has been read from a parameter file.
- make_binned_map(**kwargs)¶
- make_binned_map_splits(**kwargs)¶
- make_destriped_map(**kwargs)¶
- read_observations(**kwargs)¶
- record_profile_info(profiler: TimeProfiler)¶
- set_hwp(hwp: HWP)¶
Set the HWP to be used in the simulation
The argument must be a class derived from
HWP, for instanceIdealHWP.
- set_instrument(instrument: InstrumentInfo)¶
Set the instrument to be used in the simulation.
This function sets the
self.instrumentfield to the instance of the classInstrumentInfothat has been passed as argument. The purpose of the instrument is to provide the reference frame for the direction of each detector.Note that you should not simulate more than one instrument in the same simulation. This is enforced by the fact that if you call set_instrument twice, the second call will overwrite the instrument that was formerly set.
- set_scanning_strategy(scanning_strategy: None | ScanningStrategy = None, imo_url: str | None = None, delta_time_s: float = 60.0, append_to_report: bool = True)¶
Simulate the motion of the spacecraft in free space
This method computes the quaternions that encode the evolution of the spacecraft’s orientation in time, assuming the scanning strategy described in the parameter scanning_strategy (an object of a class derived by
ScanningStrategy; most likely, you want to useSpinningScanningStrategy). The result is saved in the member variablespin2ecliptic_quats, which is an instance of the classSpin2EclipticQuaternions.You can choose to use the imo_url parameter instead of scanning_strategy: in this case, it will be assumed that you want to simulate a nominal, spinning scanning strategy, and the object in the IMO with address imo_url (e.g.,
/releases/v1.0/satellite/scanning_parameters/) describing the parameters of the scanning strategy will be loaded. In this case, aSpinningScanningStrategyobject will be created automatically.The parameter delta_time_s specifies how often should quaternions be computed; see
ScanningStrategy.set_scanning_strategy()for more information.If the parameter append_to_report is set to
True(the default), some information about the pointings will be included in the report saved by theSimulationobject. This will be done only if the process has rank #0.
- write_healpix_map(filename: str, pixels, **kwargs) str¶
Save a Healpix map in the output folder
- Parameters:
filename (
strorpathlib.Path) – Name of the file. It must be a relative path, but it can include subdirectories.pixels – array containing the pixels, or list of arrays if you want to save several maps into the same FITS table (e.g., I, Q, U components)
- Returns:
A pathlib.Path object containing the full path of the FITS file that has been saved.
Example:
import numpy as np sim = Simulation(base_path="/storage/litebird/mysim") pixels = np.zeros(12) sim.write_healpix_map("zero_map.fits.gz", pixels)
This method saves an Healpix map into a FITS files that is written into the output folder for the simulation.
- write_observations(**kwargs)¶
- litebird_sim.simulations.get_template_file_path(filename: str | Path) Path¶
Return a Path object pointing to the full path of a template file.
Template files are used by the framework to produce automatic reports. They are produced using template files, which usually reside in the
templatessubfolder of the main repository.Given a filename (e.g.,
report_header.md), this function returns a full, absolute path to the file within thetemplatesfolder of thelitebird_simsource code.